RANJAN MANNIGE's
RESEARCH HOME FOR MOLECULAR DESIGN
Projects
Learning from molecular animals
I have explored design criteria for biological molecules (a,b, below) and artificially created molecules (c,d) using theory (e.g. network theory, statistical mechanics), math (e.g., graph theory), simulation (lattice and all-atom) and bioinformatics (structural and genomic). These studies may one day be used to help cure diseases and design novel materials at the molecular level.

News
10/2025: Prof Omar Yaghi, co-author on our experiment-theory collaboration paper [PNAS, 2015, 112(18):5591] just won the Nobel prize in Chemistry for discovering metal organic frameworks, which is a focus of the paper. This paper now has two Nobel laureates on it!
04/2018: Based on the Ramachandran number, my first Python module is published and readily downloadable via "> pip install backmap" [PyPI Link|GitHub Link].
05/2017: My fifth single-author paper revisits work by early structural biologists (Ramachandran and Miyazawa) is now out along with its citeable peer-reviews! [link|pdf].
04/2016: A nonequilibrium paper by Whitelam and I that I am very excited about is finally out in the Phys Rev E [link|pdf].
01/2016: Our review paper on Peptoid nanosheets is out [link|pdf]! Its also a journal cover [link] and one of the most downloaded paper in January '16 [link].
12/2015: My new paper on non-equilibrium assembly landscapes is out [link|pdf].
10/2015: Sir J. Fraser Stoddart, co-author on our new experiment-theory collaboration paper [PNAS, 2015, 112(18):5591] just won the Nobel prize in Chemistry for discovering mechanically interlocked molecules, which is a focus of the paper.
10/2015: My Nature paper is out [pdf]! And so is a nice writeup about it [link].
05/2015: The new PNAS paper is now featured by Phys.org!
04/2015: Our fun experiment-theory collaboration just got published at PNAS. Side note: a corresponding author won the Nobel prize a year later!
08/2014: Our beloved bilayer structure (with futuristic embellishments) is featured in the Molecular Foundry poster.
07/2014: Hot off the press: Soft Matter paper on multicomponent assembly; also, its a cover!
03/2014: Hot off the press: Proteomes hypothesis on protein origination.
03/2014: Hot off the press: Proteomes review on protein dynamism.
02/2014: Hot off the press: JCC paper on a peptoid forcefield.
10/2013: Janet Napolitano peers into peptoids.
Below are some of the projects that I have worked on:
Virus capsid assembly
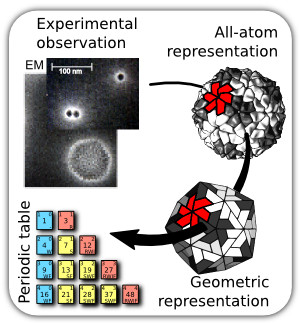
During their inter-cellular journey, viruses are protected by capsids. Capsids are protein shells mostly formed from hundreds of often chemically identical proteins called "subunits". Capsids are important to most of the virus life cycle. From a medical perspective, capsids are currently not heavily targeted for drug discovery, and our understanding of capsid assembly and disassembly could allow for the development of novel medicines. From a nanotechnology perspective, capsids are nano-scale assemblies that often assemble specifically and spontaneously; the design criteria employed by such capsids may inform the design of artificial nanoassemblies.
From graph theory and bioinformatics, my Ph.D. mentor (Charles Brooks III) and I were able to show that very complicated (and atomistically enormous) virus capsids may be simplified to canonical capsids, which are geometric entities. With this finding, we were able to obtain design criteria and constrains working on diverse viruses at the molecular (Mannige & Brooks, 2008), macromolecular (Mannige & Brooks, 2009) and evolutionary scale (Mannige & Brooks, 2010).
The final outcome of our studies, a periodic table for virus capsids, is useful in explaining diverse capsid behavior. Experimental and theoretical collaborators are encouraged to help build on this framework.
These days I am interested in how capsid proteins (subunits) are able to display such exquisite control to form specific architectures.
Origins of the first proteins (and then life!)
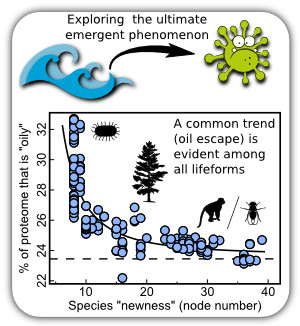
This is a question that I feel every biology-inspired scientist is eventually attracted to: how did the first functional biomolecules emerge? Understanding this question is the first step to understanding how the first lifeforms emerged.
I proposed a hypothesis (Mannige, 2014b) that oily peptides are hyper-evolvable, chemically less degradable, and so may have played an important role in the origin of the first functional proteins. The hypothesis has been validated from both simulation (Mannige, 2013a) and bioinformatics studies (Mannige, Brooks & Shakhnovich, 2012). The latter study was in collaboration with Charles Brooks III and Eugene Shakhnovich. An outcome of this project is my review on the importance of dynamism in protein structure, function and evolution: Mannige, 2014a.
Currently, I am working on the next chapter in Origins, that is how complex life emerged. Stay tuned!
Peptoid nanosheet simulation
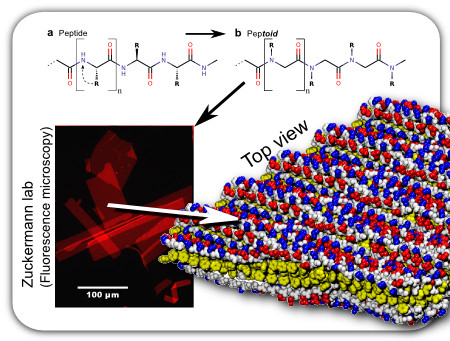
Due to one small difference from its biological counterpart (top), peptoids deviate drastically from peptides in behavior, which opens up a whole new possibility of molecular design (Mirijanian, Mannige et al., 2013). For example, some peptoids are able to assemble into nanosheets, which, while only a couple of nanometers in thickness, expand to the mesoscale in area.
Along with theorists Stephen Whitelam, Tom Haxton, Dina Mirijanian and experimentalists Ronald Zuckermann, Gloria Olivier and Caroline Proulx, we are attempting through simulation and experiment to explore the possible atom-resolution structure, assembly modes, and design criteria for these peptoid nanosheets.
Happily, the nanosheet structure that emerged from this study grabbed Janet Napolitano's interest, the once head of Homeland Security, Arizona Governor, and now the president of University of California system. To her left and right are Jeff Neaton (Director of the Molecular Foundry) and Ron Zuckermann (Facility Director, Biological Nanostructures Facility, and collaborator at the Molecular Foundry). Also happily, this story is now described in our Nature paper (see news writeup and article linked within).
MOF design criteria
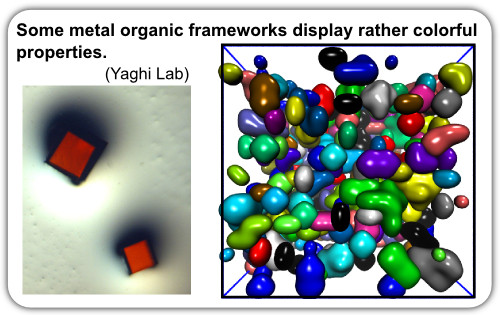
Metal Organic Frameworks (MOF) are an exciting new class of molecules with great potential utility. Currently, Steve Whitelam and I, in collaboration with experimentalists Sir J. Fraser Stoddart, Omar Yaghi, Andrew Sue, Hexiang Deng and others, are utilizing Monte Carlo simulations and graph theory to explore how two component systems can assemble into non-equilibrium frameworks but in a manner that is robust to solution conditions. This research has resulted in some interesting new modes of polycrystalline assembly (discussed in Sue, Mannigeet al. , 2015).
Learning from molecular animals
I have explored design criteria for biological molecules (a,b, below) and artificially created molecules (c,d) using theory (e.g. network theory, statistical mechanics), math (e.g., graph theory), simulation (lattice and all-atom) and bioinformatics (structural and genomic). These studies may one day be used to help cure diseases and design novel materials at the molecular level.

News
10/2025: Prof Omar Yaghi, co-author on our experiment-theory collaboration paper [PNAS, 2015, 112(18):5591] just won the Nobel prize in Chemistry for discovering metal organic frameworks, which is a focus of the paper. This paper now has two Nobel laureates on it!
04/2018: Based on the Ramachandran number, my first Python module is published and readily downloadable via "> pip install backmap" [PyPI Link|GitHub Link].
05/2017: My fifth single-author paper revisits work by early structural biologists (Ramachandran and Miyazawa) is now out along with its citeable peer-reviews! [link|pdf].
04/2016: A nonequilibrium paper by Whitelam and I that I am very excited about is finally out in the Phys Rev E [link|pdf].
01/2016: Our review paper on Peptoid nanosheets is out [link|pdf]! Its also a journal cover [link] and one of the most downloaded paper in January '16 [link].
12/2015: My new paper on non-equilibrium assembly landscapes is out [link|pdf].
10/2015: Sir J. Fraser Stoddart, co-author on our new experiment-theory collaboration paper [PNAS, 2015, 112(18):5591] just won the Nobel prize in Chemistry for discovering mechanically interlocked molecules, which is a focus of the paper.
10/2015: My Nature paper is out [pdf]! And so is a nice writeup about it [link].
05/2015: The new PNAS paper is now featured by Phys.org!
04/2015: Our fun experiment-theory collaboration just got published at PNAS. Side note: a corresponding author won the Nobel prize a year later!
08/2014: Our beloved bilayer structure (with futuristic embellishments) is featured in the Molecular Foundry poster.
07/2014: Hot off the press: Soft Matter paper on multicomponent assembly; also, its a cover!
03/2014: Hot off the press: Proteomes hypothesis on protein origination.
03/2014: Hot off the press: Proteomes review on protein dynamism.
02/2014: Hot off the press: JCC paper on a peptoid forcefield.
10/2013: Janet Napolitano peers into peptoids.
Below are some of the projects that I have worked on:
Virus capsid assembly

During their inter-cellular journey, viruses are protected by capsids. Capsids are protein shells mostly formed from hundreds of often chemically identical proteins called "subunits". Capsids are important to most of the virus life cycle. From a medical perspective, capsids are currently not heavily targeted for drug discovery, and our understanding of capsid assembly and disassembly could allow for the development of novel medicines. From a nanotechnology perspective, capsids are nano-scale assemblies that often assemble specifically and spontaneously; the design criteria employed by such capsids may inform the design of artificial nanoassemblies.
From graph theory and bioinformatics, my Ph.D. mentor (Charles Brooks III) and I were able to show that very complicated (and atomistically enormous) virus capsids may be simplified to canonical capsids, which are geometric entities. With this finding, we were able to obtain design criteria and constrains working on diverse viruses at the molecular (Mannige & Brooks, 2008), macromolecular (Mannige & Brooks, 2009) and evolutionary scale (Mannige & Brooks, 2010).
The final outcome of our studies, a periodic table for virus capsids, is useful in explaining diverse capsid behavior. Experimental and theoretical collaborators are encouraged to help build on this framework.
These days I am interested in how capsid proteins (subunits) are able to display such exquisite control to form specific architectures.
Origins of the first proteins (and then life!)

This is a question that I feel every biology-inspired scientist is eventually attracted to: how did the first functional biomolecules emerge? Understanding this question is the first step to understanding how the first lifeforms emerged.
I proposed a hypothesis (Mannige, 2014b) that oily peptides are hyper-evolvable, chemically less degradable, and so may have played an important role in the origin of the first functional proteins. The hypothesis has been validated from both simulation (Mannige, 2013a) and bioinformatics studies (Mannige, Brooks & Shakhnovich, 2012). The latter study was in collaboration with Charles Brooks III and Eugene Shakhnovich. An outcome of this project is my review on the importance of dynamism in protein structure, function and evolution: Mannige, 2014a.
Currently, I am working on the next chapter in Origins, that is how complex life emerged. Stay tuned!
Peptoid nanosheet simulation

Due to one small difference from its biological counterpart (top), peptoids deviate drastically from peptides in behavior, which opens up a whole new possibility of molecular design (Mirijanian, Mannige et al., 2013). For example, some peptoids are able to assemble into nanosheets, which, while only a couple of nanometers in thickness, expand to the mesoscale in area.
Along with theorists Stephen Whitelam, Tom Haxton, Dina Mirijanian and experimentalists Ronald Zuckermann, Gloria Olivier and Caroline Proulx, we are attempting through simulation and experiment to explore the possible atom-resolution structure, assembly modes, and design criteria for these peptoid nanosheets.
Happily, the nanosheet structure that emerged from this study grabbed Janet Napolitano's interest, the once head of Homeland Security, Arizona Governor, and now the president of University of California system. To her left and right are Jeff Neaton (Director of the Molecular Foundry) and Ron Zuckermann (Facility Director, Biological Nanostructures Facility, and collaborator at the Molecular Foundry). Also happily, this story is now described in our Nature paper (see news writeup and article linked within).
MOF design criteria

Metal Organic Frameworks (MOF) are an exciting new class of molecules with great potential utility. Currently, Steve Whitelam and I, in collaboration with experimentalists Sir J. Fraser Stoddart, Omar Yaghi, Andrew Sue, Hexiang Deng and others, are utilizing Monte Carlo simulations and graph theory to explore how two component systems can assemble into non-equilibrium frameworks but in a manner that is robust to solution conditions. This research has resulted in some interesting new modes of polycrystalline assembly (discussed in Sue, Mannige
© Ranjan Mannige